Welcome to the Dr. Gillies Lab
Latest News
November 17th, 2021
2021 Cancer Biology Outstanding Research Award Recipients
This annual award recognizes students in the Cancer Biology PhD Program who demonstrate an exceptional level of performance in research as evidenced by mentor evaluations, conference presentations, and publications, etc. We are delighted to announce that members of the award selection committee have chosen Carly Harro and Bryce Ordway as the recipients of this award for 2021.
Congratulations!


August 4th, 2021
The American Association for Cancer Research (AACR) recently announced the formation of its seventh and newest scientific working group, which will be dedicated to supporting cancer evolution research.
Moffitt's Drs. Robert Gatenby, Robert Gillies, and Andriy Marusyk were named members of the AACR Cancer Evolution Working Group Executive Committee.
The mission of the AACR Cancer Evolution Working Group is to advance cancer diagnostic, therapeutic, and prevention strategies by fostering a fundamental understanding of cancer evolution amongst its members and the broader cancer research community. Membership in the AACR Cancer Evolution Working Group is free and open to all members of the AACR interested in cancer evolution.



January 13th, 2021
Moffitt Postdoc Receives 2021 Laurence P. Clarke Young Scientist Award in Quantitative Imaging

Dr. Wei Mu, a PhD graduate from the University of Chinese Academy of Science in Beijing, has been a postdoctoral fellow at the Moffitt Cancer Center for the past four years.
Her research interests involve the development of machine learning models to analyze multimodal medical images for the early diagnosis of cancer and to aid in the decision of individualized treatment planning. Her dissertation pioneered work in automatic segmentation methods and texture analysis of FDG PET images to improve prognostic models for both cervical and colorectal cancers. She continues her work in PET image analysis at Moffitt Cancer Center by developing additional PET/CT radiomics features through convolution of PET and CT images.
Dr. Mu has received several awards for her work over the last few years. She is the recipient of the Outstanding Student and Graduate Award from her undergraduate school and received several scholarships to the University of Chinese academy of Science. She was one of four finalists in the 2020 WMIS Young Investigator Award and recipient of the Women in Molecular Imaging Network Scholar Award from WMIS. She is an author on over a dozen peer-reviewed articles.
Grant Updates
October 13th, 2021
Drs. Damaghi and Silva Awarded NCI U01
Drs. Mehdi Damaghi and Ariosto Silva (MPI) were awarded an NCI U01 of $3,417,517 over 5 years for their research project titled "Ecology and Evolution of Breast Carcinogenesis."
Drs. Marilyn Bui, Brian Czerniecki, Robert Gatenby, and Robert Gillies will serve as Co-Investigators on the project.
Congratulations, Drs. Damaghi and Silva!


July 14th, 2021
Dr. Schabath Awarded FBRP/Bankhead-Coley Funds
Dr. Matt Schabath was awarded a Florida Biomedical Research Program (FBRP)/Bankhead-Coley grant of $1,327,180 over 5 years for his research project titled “Non-invasive radiomic biomarkers to predict treatment response for immunotherapy of lung cancer."
Drs. Robert Gillies, Issam El Naqa, and Jhanelle Gray will serve as Co-Investigators on the project.
Congratulations, Dr. Schabath!
April 14th, 2021
Drs. Gillies and Niell Awarded NCI R01
Drs. Bob Gillies and Bethany Niell were awarded an NCI R01 of $3,430,507 over 5 years for their research project titled “Radiomics and Pathomics to Predict Upstaging of DCIS."
Co-Investigators on the project are Drs. Dana Ataya, Marilyn Bui, and Brian Czerniecki.
Congratulations, Drs. Gillies and Niell!


March 11th, 2020
Drs. Gillies and Pilon-Thomas Awarded NIH/NCI R01
Drs. Bob Gillies and Shari Pilon-Thomas were recently awarded an NIH/NCI R01 for $1,878,475 over five years for a research project titled “Imaging Acidosis and Immune Therapy in PDAC." This is Dr. Pilon-Thomas's first R01. Congratulations!
CO-Is on the project are Drs. Arig Ibrahim Hashim, Barbara Centeno, and Jason Fleming.
Congratulations, Drs. Gillies and Pilon-Thomas!


Robert J. Gillies, PhD.
Robert J. Gillies and his lab are focused on understanding cancers as complex, heterogeneous and dynamic systems. Along with his long-time collaborator, Robert A. Gatenby, they share a core belief that, due to genomic plasticity and microenvironmental heterogeneity, cancers can only be understood through the lens of Darwinian Evolution. Dr. Gillies is an experimentalist whose work spans molecular, cellular, animal models, and image analytics. He is the Martin Silberger Chair of Cancer Physiology, Director of the Center of Excellence in Cancer Imaging and Technology, and Scientific Director of the Small Animal Imaging Lab, SAIL.
May 2017, Cancer Research cover, from Arig Ibrahim-Hashim, Robert J. Gillies et al. "Defining Cancer Subpopulations by Adaptive Strategies Rather Than Molecular Properties Provides Novel Insights into Intratumoral Evolution"
Gillies Lab at the Moffitt PSOC Site Visit
Publication Highlights:
Moffitt Researchers Develop Non-invasive Approach to Measure Biomarker Levels, Predict Outcomes in Lung Cancer Patients published in Journal for Immunotherapy of Cancer
Dr. Wei Mu
June 9, 2021 (Moffitt MRI Signals June 23, 2021)
Tests that analyze biomarkers are used during cancer management to guide treatment and provide information about patient prognosis. These tests are often performed on tissue biopsy samples that require invasive procedures and can lead to significant side effects. In a new article published in the Journal for ImmunoTherapy of Cancer, Moffitt Cancer Center researchers show that PET/CT images can be used to measure levels of the PD-L1 biomarker of non-small cell lung cancer (NSCLC) patients in a non-invasive manner and, in turn, predict a patient’s response to therapy.
Moffitt Researchers Develop Model to Predict Non-Small Cell Lung Cancer Patient Outcomes to Immunotherapy published in JNCI Cancer Spectrum
Dr. Ilke Tunali
May 13, 2021 (Moffitt MRI Signals August 25, 2021)
 Immunotherapy agents that inhibit the molecules PD1, PD-L1 or CTLA-4 have become widely used in clinical practice to treat non-small cell lung cancer, or NSCLC. Approximately 20% to 50% of advanced NSCLC patients have strong responses to immunotherapy and show prolonged survival, but the remaining patients often have poor responses. There is an urgent need to identify biomarkers that can predict which patients will not respond to therapy to avoid unnecessary treatment and administer potential beneficial drugs instead.
Immunotherapy agents that inhibit the molecules PD1, PD-L1 or CTLA-4 have become widely used in clinical practice to treat non-small cell lung cancer, or NSCLC. Approximately 20% to 50% of advanced NSCLC patients have strong responses to immunotherapy and show prolonged survival, but the remaining patients often have poor responses. There is an urgent need to identify biomarkers that can predict which patients will not respond to therapy to avoid unnecessary treatment and administer potential beneficial drugs instead.
Moffitt Research published in European Journal of Nuclear Medicine and Molecular Imaging
Dr. Narges Tafreshi
March 26, 2021 (Moffitt MRI Signals April 7, 2021)
A research article titled "Preclinical evaluation of [225 Ac]Ac-DOTA-TATE for treatment of lung neuroendocrine neoplasms" was recently published in European Journal of Nuclear Medicine and Molecular Imaging.
There is significant interest in the development of targeted alpha-particle therapies (TATs) for the treatment of solid tumors. The metal chelator-peptide conjugate, DOTA-TATE, loaded with the β-particle emitting radionuclide 177Lu ([177Lu]Lu-DOTA-TATE) is now standard care for neuroendocrine tumors that express the somatostatin receptor 2 (SSTR2) target. A recent clinical study demonstrated efficacy of the corresponding [225Ac]Ac-DOTA-TATE in patients that were refractory to [177Lu]Lu-DOTA-TATE. Herein, investigators report the radiosynthesis, toxicity, biodistribution, radiation dosimetry, and efficacy of [225Ac]Ac-DOTA-TATE in small animal models of lung neuroendocrine neoplasms (NENs).
Results from the study suggest significant potential for the clinical translation of [225Ac]Ac-DOTA-TATE for lung NENs.
Deep-learning and MR images to target hypoxic habitats with evofosfamide in preclinical models of sarcoma published in Theranostics
Dr. Bruna V. Jardim-Perassi
March 11, 2021
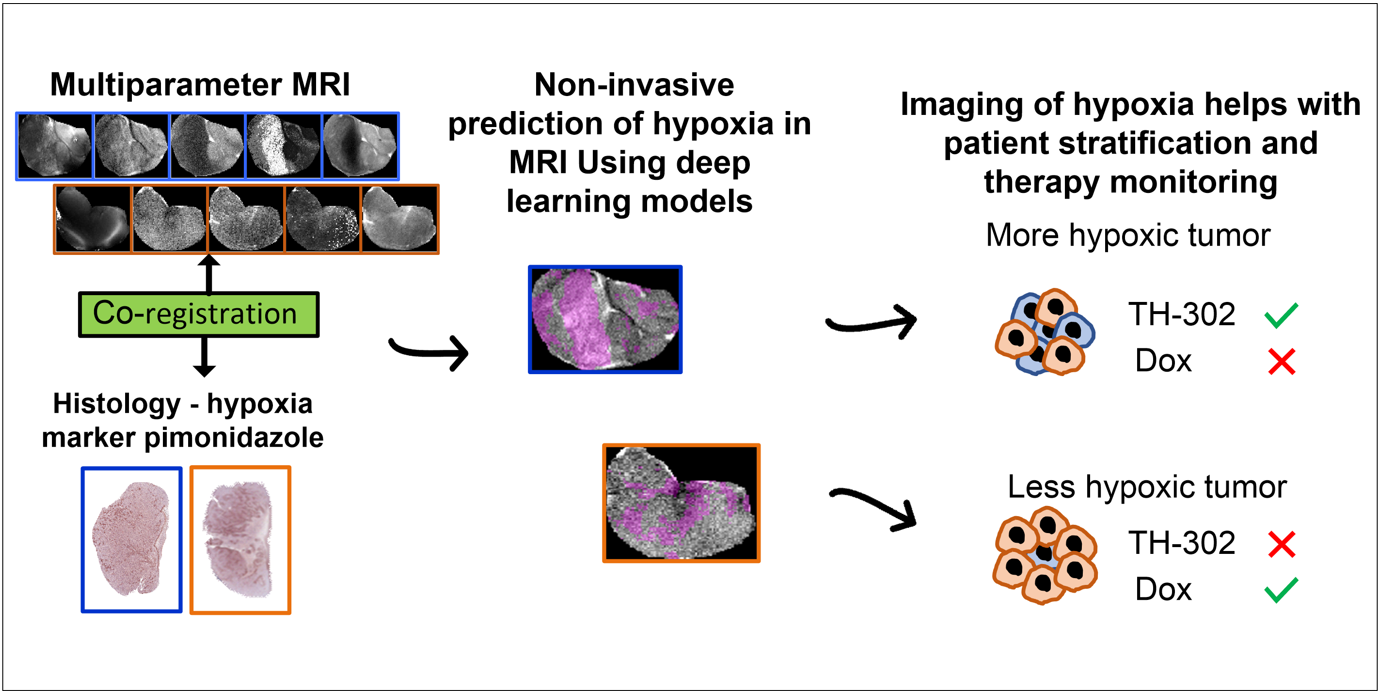
 In this paper, we developed deep-learning models based on multiparametric MRI co-registered with pimonidazole-stained histology to predict hypoxia in MRI. Using these models, we found that hypoxia measurement prior to therapy can non-invasively presage the response to the hypoxia-activated prodrug evofosfamide (TH-302) and longitudinally monitor anti-tumor effects in pre-clinical models of sarcoma.
In this paper, we developed deep-learning models based on multiparametric MRI co-registered with pimonidazole-stained histology to predict hypoxia in MRI. Using these models, we found that hypoxia measurement prior to therapy can non-invasively presage the response to the hypoxia-activated prodrug evofosfamide (TH-302) and longitudinally monitor anti-tumor effects in pre-clinical models of sarcoma.
Moffitt Researchers Identify How Cancer Cells Adapt to Survive Harsh Tumor Microenvironments published in PNAS
Dr. Mehdi Damaghi
January 19, 2021 (Moffitt MRI Signals January 20, 2021)
Cells need energy to survive and thrive. Generally, if oxygen is available, cells will oxidize glucose to carbon dioxide, which is very efficient, much like burning gasoline in your car. However, even in the presence of adequate oxygen, many malignant cells choose instead to ferment glucose to lactic acid, which is a much less efficient process. This metabolic adaptation is referred to as the Warburg Effect, as it was first described by Otto Warburg almost a century ago. Ever since, the conditions that would evolutionarily select for cells to exhibit a Warburg Effect have been in debate, as it is much less efficient and produces toxic waste products.
“The Warburg Effect is misunderstood because it doesn’t make sense that a cell would ferment glucose when it could get much more energy by oxidizing it. Our current study goes to the heart of this problem by defining the microenvironmental conditions that exist in early cancers that would select for a Warburg phenotype. This is important because such cells are much more aggressive and likely to lead to cancers that are lethal,” said Mehdi Damaghi, Ph.D., study first author and research scientist in the Cancer Physiology Department at Moffitt Cancer Center.
Artificial selection for host resistance to tumour growth and subsequent cancer cell adaptations: an evolutionary arms race published in British Journal of Cancer
Dr. Arig Ibrahim-Hashim
January 19, 2021 (Nature January 19, 2021)
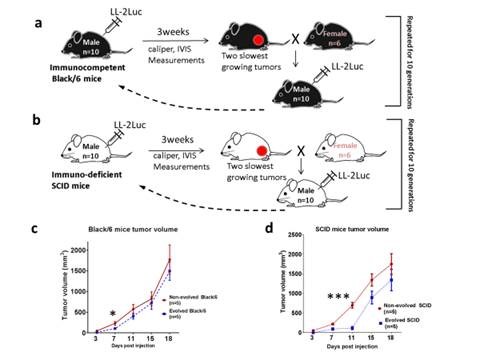
In this publication, we have bred a line of SCID and Black/6 mice that are more resistant to cancer malignancy than their wild type relatives using a subcutaneous inoculation paradigm with LL/2-Luc-M38 cancer cells. Our results show that the evolved phenotype results from different mechanisms in each strain. In the SCID mice, suppression of tumor growth was through the change in the biology of the stromal cells, most likely the skin fibroblasts, and in Black/6 mice is through the increase in the immune-mediated killing of cancer cells.
Heterogeneity analysis of MRI T2 maps for measurement of early tumor response to radiotherapy published in NMR in Biomedicine
Dr. Michal Tomaszewski
December 15, 2020 (Moffitt MRI Signals January 6, 2021)
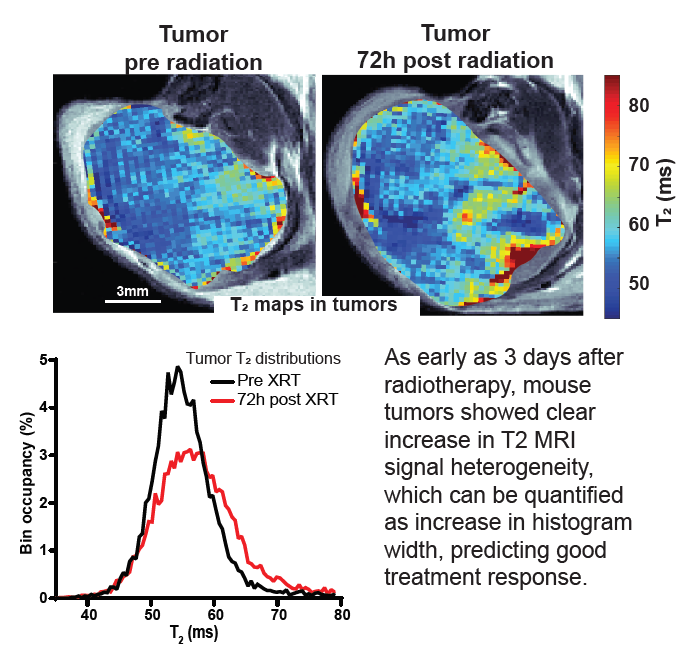
Response to radiotherapy is a complex function of multiple factors, and patients vary significantly in the extent to which irradiation helps slow the development of their disease. This is particularly important in pancreatic cancer, where short progression times mean that timely and effective treatment is crucial. In this paper we show in mouse model of pancreatic cancer, that analysis of T2 mapping in Magnetic Resonance Imaging coupled with appropriate analysis can be used to capture the early signs that relate to the degree of radiotherapy response later on. Our method, validated against histological data, allows to quantify the amount of inhomogeneity (“heterogeneity”) in the images which relates to the early signs of the desired harmful effect of the radiation on tumor tissue. In the future this method can be tested clinically and ultimately used to help personalize radiotherapy for pancreatic cancer patients.
Mutation-selection balance and compensatory mechanisms in tumour evolution published in Nature Reviews Genetics
Dr. Erez Persi
November 30, 2020
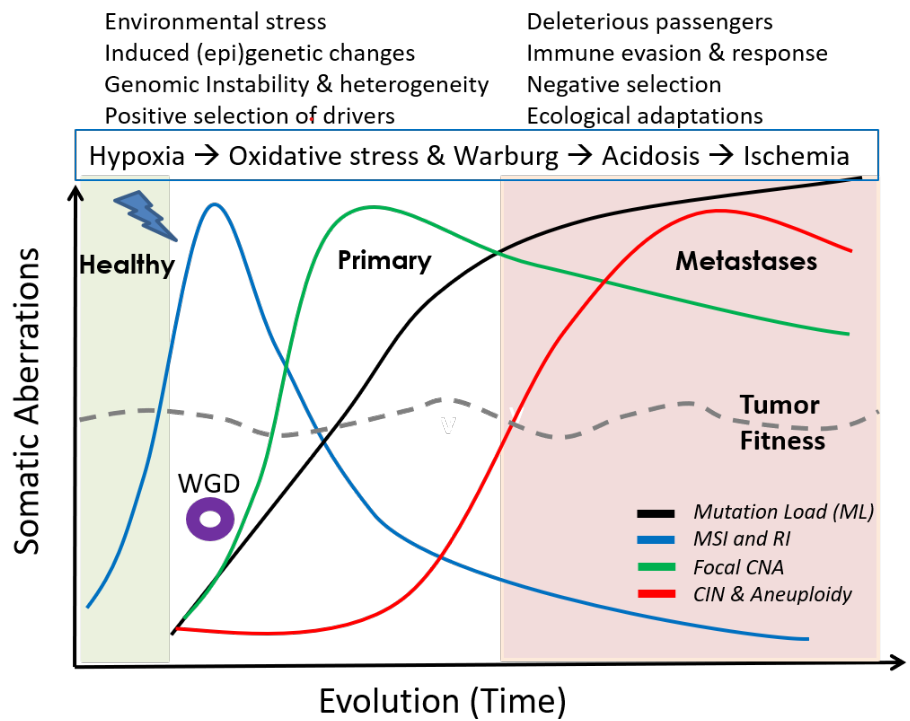
A synthesis of mutational and clinical patient data, combined with literature from different independent laboratories, provides insights into the rich, temporal dynamics and compensatory roles of different genomic aberrations in tumor evolution, and their relation to phenotypic plasticity; explaining how cancers remain fit over time. A small step towards the general rules of cancer evolution, and a road map for cancer early detection and development of smart therapeutics.
Non-invasive decision support for NSCLC treatment using PET/CT radiomics published in Nature Communications
Dr. Wei Mu
October 16, 2020 (Moffitt MRI Signals October 21, 2020)
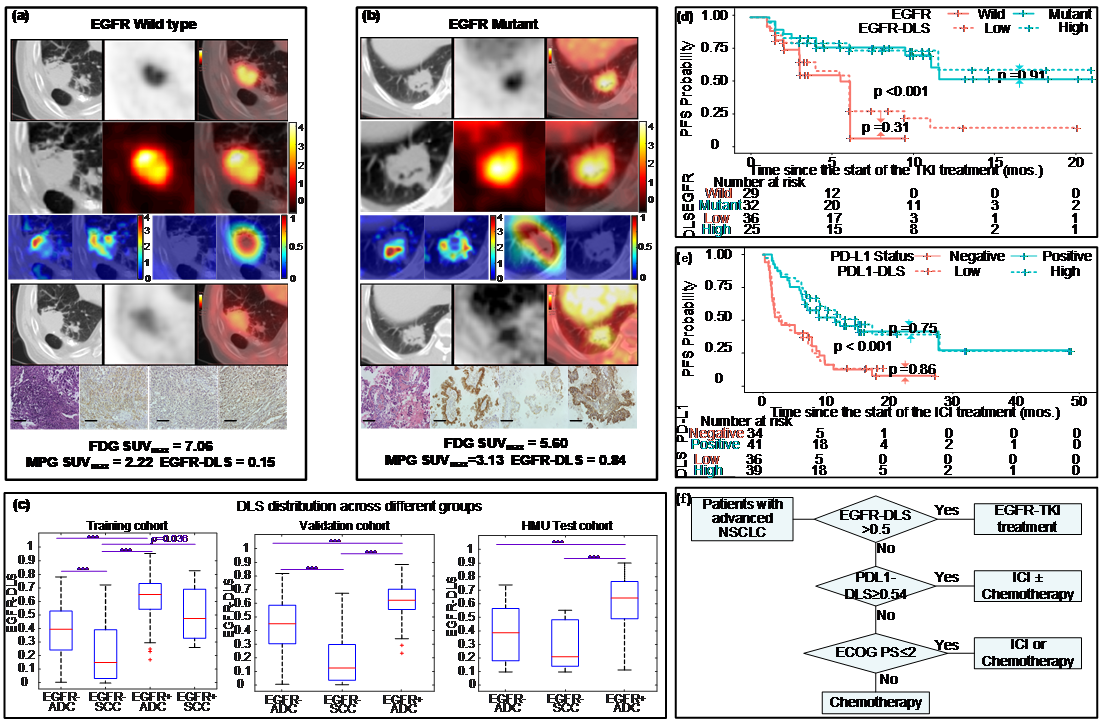
EGFR, or epidermal growth factor receptor, is a common mutation found in non-small cell lung cancer patients, and can be a predictor for treatment. However, the current biopsy-based detection techniques are invasive, and may fail to yield actionable results due to insufficient quantity or quality of the tissue. In this work, we developed a deep learning model to predict EGFR mutation using 18F-FDG PET/CT images since it is widely used and the glucose radiotracer used could be affected by EGFR activation and inflammation. Further, we found that the EGFR deep learning score was positively associated with longer progression free survival in patients treated with tyrosine kinase inhibitors (TKI), and negatively associated with durable clinical benefit and longer progression free survival in patients being treated with immune checkpoint inhibitor (ICI). Therefore, together with our previously developed PD-L1 deep learning prediction model using PET/CT images, we developed a non-invasive treatment decision support to make treatment plan.
Radiomics of (18)F-FDG PET/CT images predicts clinical benefit of advanced NSCLC patients to checkpoint blockade immunotherapy published in European Journal of Nuclear Medicine and Molecular Imaging
Dr. Wei Mu
December 5, 2019 (Moffitt MRI Signals January 8, 2020)
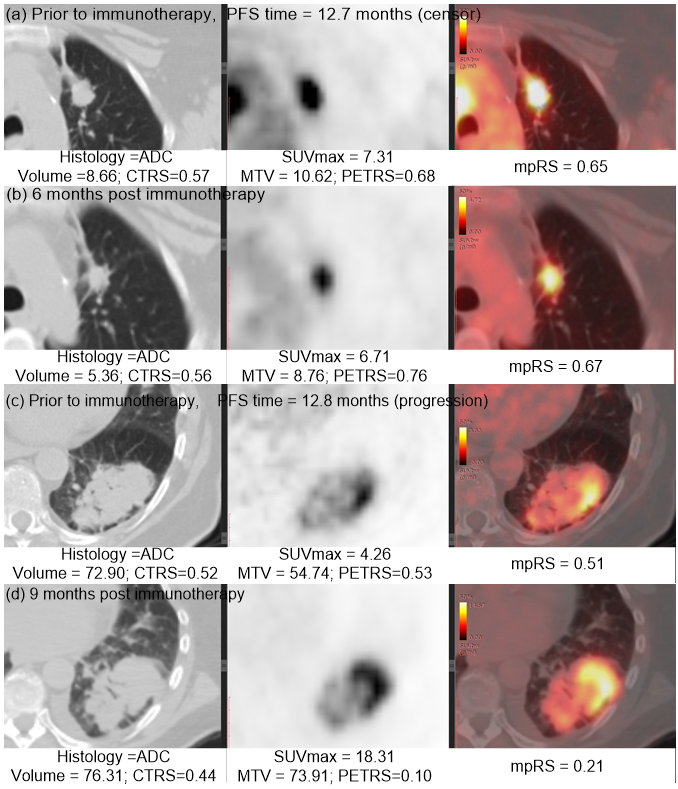
Immunotherapy has improved outcomes for patients with non-small cell lung cancer (NSCLC), yet durable clinical benefit (DCB) is experienced in only a fraction of patients. Therefore, we developed an effective and stable radiomics signature by the combination of PET, CT and the fusion Kullback–Leibler Divergence features, which may serve as a predictive biomarker for immunotherapy response. This signature could be used prior to initiation of immunotherapy to identify NSCLC patients most likely to benefit from immunotherapy, and could also be leveraged to improve the non-invasive treatment decision support in the treatment of advanced NSCLC patients.
Systems analysis of intracellular pH vulnerabilities for cancer therapy published in Nature Communications
Drs. Damaghi, Gillies, Cleveland & Colleagues
July 31, 2018 (Moffitt MRI Signals August 15, 2018)
Research Scientist, Mehdi Damaghi, in Gillies lab was co-lead author a paper recently published in Nature Communications that utilized a systems analysis of intracellular pH vulnerabilities for cancer therapy. The team, which included Dr. Bob Gillies, and John Cleveland, developed a computational methodology that explores how intracellular pH (pHi) can modulate metabolism. Experimental testing of the novel strategy revealed that it is particularly effective against aggressive phenotypes. The study suggests essential roles of pHi in cancer metabolism and provides a conceptual and computational framework for exploring pHi roles in other biomedical domains. Moffitt's Molecular Genomics and Analytical Microscopy cores were also utilized.
Eco-evolutionary Causes and Consequences of Temporal Changes in Intramural Blood Flow published in Nature Reviews Cancer
Drs. Gillies, Brown, Anderson, and Gatenby
June 11, 2018 (Moffitt MRI Signals July 18, 2018)
Their opinion posits that temporal changes in blood flow are commonly observed in malignant tumors, but the evolutionary causes and consequences are rarely considered. The authors propose that stochastic temporal variations in blood flow and microenvironmental conditions arise from the eco-evolutionary dynamics of tumor angiogenesis in which cancer cells, as individual units of selection, can influence and respond only to local environmental conditions.
Temporal variations in intratumoural blood flow, which occur through the promotion of cancer cell phenotypes that facilitate both metastatic spread and resistance to therapy, may have substantial clinical consequences.
Quantitative Imaging (QI)
Quantitative Imaging is an indispensable approach to study intratumoral heterogeneity. Dr. Gillies' group develops and applies Image Analytics to histological and radiological images to investigate the spatial relationships of cells and microenvironments in clinical and pro-clinical tumors.
Projects include:
Tumor pH
Solid tumors are acidic due to elevated glycolysis combined with poor perfusion. Dr. Gillies has developed methods to image tumor pH, and study the causes and consequence of this acidity. Recent focus has been on developing methods to interfere with tumor acidosis for improving therapy in the clinic.
Projects include:
Physical Sciences in Oncology Center at Moffitt Cancer Center
Upcoming Events
- To Be Announced